Choosing NEPA21 vs Viral Delivery in Brain Organoids
How labs decide based on organoid state, lumen access, cargo size, and mosaic vs stable expression.
Brain organoids are thick, regionally patterned 3D tissues, so delivery strategy matters early. In practice, the best method usually depends on four things: organoid stage, access to ventricular-like lumens, cargo size, and whether you want mosaic or broad expression.
For many labs, the key choice is:
NEPA21 for fast, flexible, non-viral delivery
vs
Viral delivery for longer-term, more uniform expression
Navigation Shortcuts
The 30-second answer
Choose NEPA21 when you need:
- fast validation
- lumen-targeted delivery
- large cargo such as plasmids or CRISPR RNPs
- mosaic labelling or cell-autonomous phenotypes
Choose viral delivery when you need:
- stable expression over weeks to months
- broader labelling in thicker or more mature tissue
- long-term functional assays
- pooled or selection-based workflows
A practical workflow many labs use:
NEPA21 first for fast construct or guide validation → viral delivery later for stable assays
(Fast check)
| 1. | Do you need stable expression for weeks to months, lineage tracing, or pooled screens? |
| → Yes: Viral (usually lenti; AAV depending on target) → No / transient is fine: NEPA21 |
| 2. | Is the organoid early or small and lumen-accessible (< ~500 µm, thin ECM)? |
| → Yes: NEPA21 → it is thicker or more mature (> ~1 mm): Viral delivery is usually more practical |
| 3. | Is your cargo large or difficult to package into virus, such as CRISPR RNPs or big plasmids > 8 kb? |
| → Yes: NEPA21 → No: Either can work; decide based on duration and labelling pattern |
| 4. | Do you want mosaic, cell-autonomous phenotypes, or sparse labelling? |
| → Yes: NEPA21 → No: you want broad uniform labelling/expression: Viral |
NEPA21 vs Viral Delivery At a Glance
| Criterion | NEPA21 Electroporation | Viral Delivery (Lentivirus / AAV) |
|---|---|---|
| Expression duration | Typically transient, days to weeks | Longer-term; often stable with lentivirus, episomal with AAV |
| Cargo flexibility | DNA, mRNA, RNPs; large constructs feasible | More packaging constraints; AAV especially size-limited |
| Best stage | Early organoids, rosettes, lumen-accessible tissue | Mid-to-late organoids, thicker tissue |
| Labelling pattern | Mosaic, tuneable, sparse-friendly | Broader, more uniform (MOI/tropism dependent) |
| Throughput / cost | Low cost, fast turnaround (often same-day), no viral production | Slower setup and expression timeline |
| Biosafety | Minimal (non-viral) | Typically BSL-2 handling |
| Tissue access bias | Strong for lumen-facing/apical regions | Better suited to broader tissue exposure |
Why labs choose NEPA21 in brain organoids
| NEPA21 is commonly chosen when labs need a delivery method that is fast, flexible, and well suited to pilot-stage engineering in fragile epithelial 3D systems. Typical advantages labs cite: |
|
Method snapshots
NEPA21 electroporation in brain organoids
When labs choose it:
- Early neural induction / neuroepithelial cyst stages (often ~day 15–30).
- Mosaic, cell-autonomous phenotypes (polarity, migration, differentiation).
- Large constructs or CRISPR RNP delivery.
- Lumen targeting (“apical electroporation” analog to in utero electroporation).
Typical strengths:
- Fast setup; rapid go/no-go results.
- Strong expression in lumen-facing progenitors (when access is good).
- No integration footprint.
- Mosaicism supports within-organoid comparisons.
Typical trade-offs:
- Efficiency drops with organoid age and ECM/thickness.
- Transient expression; plasmids dilute with divisions.
- Heterogeneous depth (apical >> deep neurons).
- Late-stage organoids can be fragile if parameters aren’t optimized.
Typical workflow:
- Prepare early organoids (often pre-embed or briefly handle to allow access).
- Microinject DNA/RNP mix into lumen/rosette space.
- Apply NEPA21 poring + transfer pulses (optimize per stage/electrode/tissue).
- Recovery culture (often ROCK inhibitor) and re-embed if needed.
- Analyse ~3–10 days for transient assays (or longer depending on readout).
Viral delivery (Lenti or AAV)
When labs choose it:
- Mature organoids (often ~50–200 days, depending on model).
- Need stable labelling (GCaMP, synaptic reporters, tracing).
- Pooled CRISPRi/a, barcode lineage screens.
- Even labelling across multiple layers (as feasible).
Typical strengths:
- Gentle on tissue (no electroporation shock).
- Stable long-term expression (especially lenti).
- Can select/sort infected cells (depending on design).
- AAV serotype choice can bias targeting by cell type/layer.
Typical trade-offs:
- Vector production/purchase adds time (often 1–2+ weeks).
- Size limits (AAV ~4.7 kb; lenti ~8–9 kb typical).
- Biosafety requirements (often BSL-2).
- MOI can influence transcriptional state; requires controls.
Typical workflow:
- Obtain titered viral stocks.
- Incubate intact organoids or slices in virus (sometimes with agitation/spinoculation).
- Allow expression to develop (AAV often days; lenti often ≥7 days for stable readouts).
- Optional selection/sorting for uniformity.
- Proceed to functional assays (imaging, electrophysiology, tracing, screens).
Typical use cases you can map to your goal
| Goal | Common choice | Why |
|---|---|---|
| Quick promoter or cDNA function test | NEPA21 plasmid | Fast, transient readout |
| CRISPR knockout using Cas9 RNPs | NEPA21 | Large cargo; mosaic KO for cell-autonomous effects |
| Stable reporter (GCaMP, synapsin-GFP) | Lentivirus | Long-term expression for functional assays |
| Inducible CRISPRi/a or pooled screens | Lentivirus | Stable integration and selection |
| Connectivity mapping / tracing | AAV (serotype-specific) | Tropism + longer assays |
State-based rule of thumb
| Stage / Model | Common choice | Why |
|---|---|---|
| Neuroepithelial cysts (day ~10–20) | NEPA21 | Lumen accessible; fast; IUE-like targeting |
| Early cortical plate–like (day ~30–60) | Either (depends on size) | Smaller organoids tolerate NEPA21; viral helps uniformity/stability |
| Large or fragile intact organoids | Viral | Less mechanical stress from electroporation workflows |
| Mature cortical organoids (day > ~90) | Viral | Thick tissue; higher electroporation risk; long-term functional assays |
| Slice cultures / fused assembloids | Viral | More even spread across layers; regional targeting possible |
Published examples in brain/cortical organoid-related workflows
Examples where NEPA21 is used directly in cortical or brain organoids after microinjection or in chamber/cuvette formats.Taniguchi-Ikeda et al., iScience (2021)
FCMD brain organoid migration assay
- What they delivered: plasmid
- Model/stage: cortical organoids; microinjection into ventricle/rosette regions
- Hardware: “electroporation glass chamber” filled with Opti-MEM; plate electrodes in chamber (model not specified)
- NEPA21 use-point: microinject → whole-organoid electroporation
- Notable detail: reported pulse conditions: 5 pulses, 125 V, 50 ms, 1 s intervals
Buijsen et al., Biomedicines (2024)
ASO delivery in 2D/3D hiPSC-derived neuronal models
- What they delivered: antisense oligonucleotides (ASOs)
- Model/stage: hiPSC-derived cortical organoids
- Hardware: electroporation cuvette placed into CU500 Cuvette Chamber; also mentions a “NEPA electrode” (model not specified)
- NEPA21 use-point: organoid electroporation (cuvette format)
Hendriks et al., Cell (2024)
Human fetal brain self-organizes into long-term expanding organoids
Long-term expanding fetal brain organoids
- What they delivered: not listed here (method focus is the platform)
- Model/stage: fetal brain organoid pieces
- Hardware: 2 mm gap cuvette
- NEPA21 use-point: whole-organoid/piece electroporation in cuvette format
Yamauchi et al., bioRxiv (posted 2025)
Heterochronic scaling of neurogenesis for species-specific dosing of cortical excitatory subtypes
Cortex-related workflow
- Hardware: CUY650P5 tweezer electrodes (5 mm platinum disk)
- NEPA21 use-point: tissue-level electroporation with tweezer format
NEPA21 used upstream to engineer cells that then form organoids
Examples where NEPA21 is used to edit hPSCs or iPSCs first, followed by organoid generation.
Pagliaro et al., Nature Communications (2023)
ENO/cortical identity induction workflow
- What they did: electroporate hESCs used downstream for cortical organoid work
- Hardware: 2 mm gap cuvettes (EC-002S)
- NEPA21 use-point: upstream cell engineering step
Kim et al., Molecular Cells (2022)
Aberrant Cortical Layer Development of Brain Organoids Derived from Noonan Syndrome-iPSCs
Noonan syndrome iPSC-derived brain organoids
- What they did: gene-editing step for iPSCs used to generate cortical organoids
- Hardware: cuvettes with NEPA21 (gap/model not specified)
- NEPA21 use-point: upstream cell editing
Hong et al., Bioengineering & Translational Medicine (2024)
AAVS1-targeted stable ChR2 expression for brain organoids
- What they did: engineer hPSCs for forebrain organoids
- Hardware: 1 mm gap cuvette; also lists CUY650P5 alongside NEPA21
- NEPA21 use-point: upstream engineering for consistent downstream optogenetics
Kim et al., Science Advances (2025)
Perturbed cell fate decision by schizophrenia-associated AS3MTd2d3 isoform during corticogenesis
Corticogenesis/schizophrenia-associated isoform study
- As listed: NEPA21 + 2 mm gap cuvettes (EC-002S)
- NEPA21 use-point: upstream engineering supporting downstream organoid experiments
Decision Flowchart: Choosing NEPA21 vs. Viral Delivery in Brain Organoid Experiments
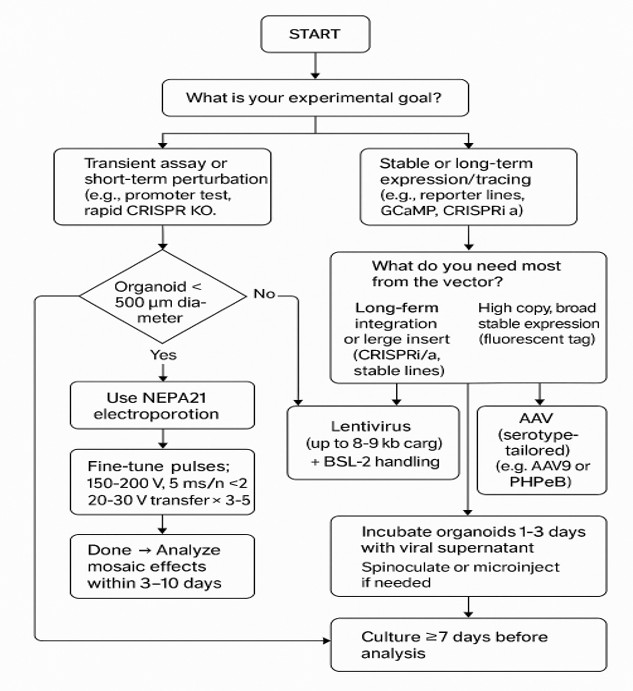
How to choose for your experiment
Choose NEPA21 when your workflow is early-stage, fast-turnaround, cargo-heavy, or benefits from mosaic readouts.NEPA21 for rapid validation → viral delivery for stable downstream assays.
Talk to us about your organoid workflow
Share your:
- organoid size or culture state
- cargo type
- desired expression pattern
- readout timeline
We can help recommend:
- the best delivery approach
- electrode format
- an optimization strategy aligned to your assay